Eric Galburt
Welcome Bridget Lu!
Washington University undergraduate Bridget Lu has begun working in the laboratory while she works towards her degree.
Welcome Eun Sil and Vincent!
BBSB students Eun Sil Rollins and Vincent Nicholson have begun their rotations in the lab. We’re glad they’re here!
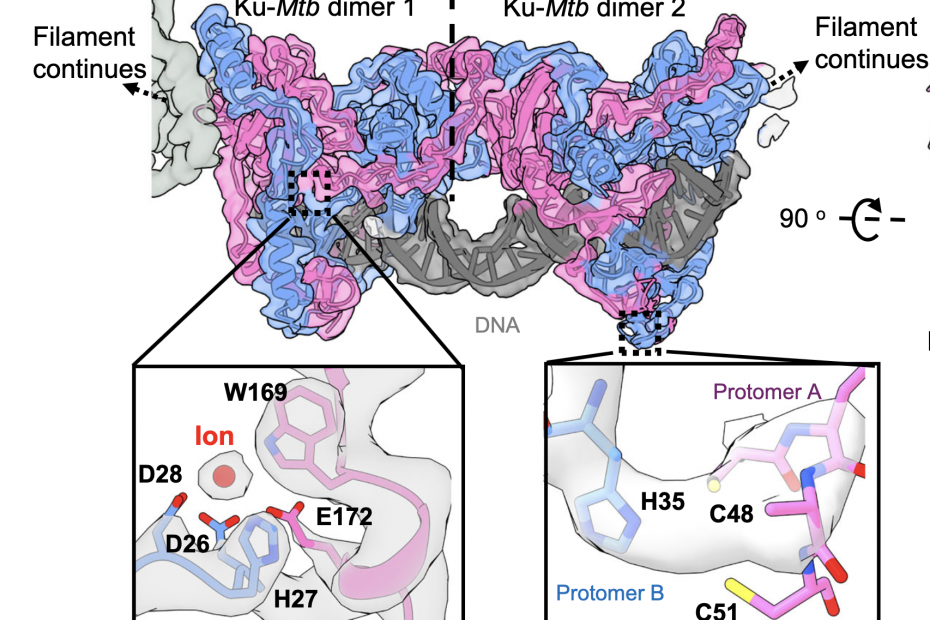
The structure of DNA repair factor Ku from Mycobacterium tuberculosis!
Now published cryo-EM work from our collaborators in Amanda Chaplin‘s lab (Leicester University, UK) has revealed the structure of a bacterial Ku for the first time and provides insight into the mechanisms of non-homologous end-joining of double-stranded breaks in this… Read More »The structure of DNA repair factor Ku from Mycobacterium tuberculosis!
Congratulations to Audrey on passing her QE!
Audrey passed her Qualifying Exam with high marks! Well done Audrey!